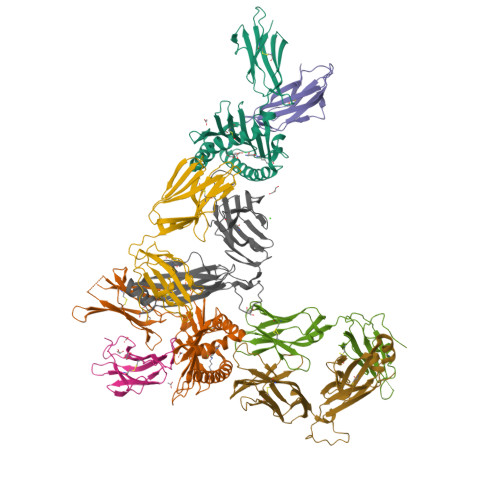
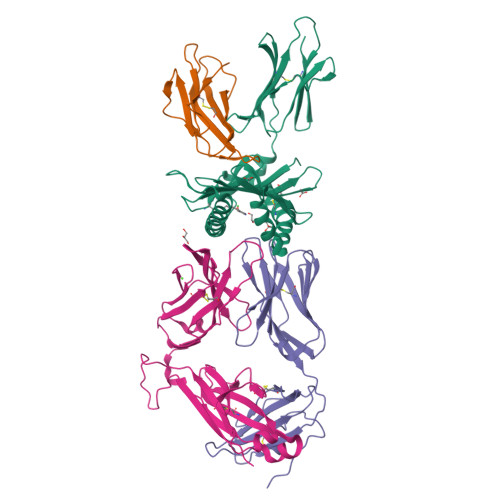
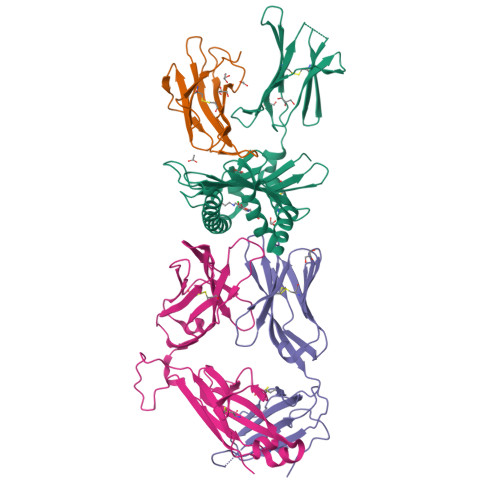
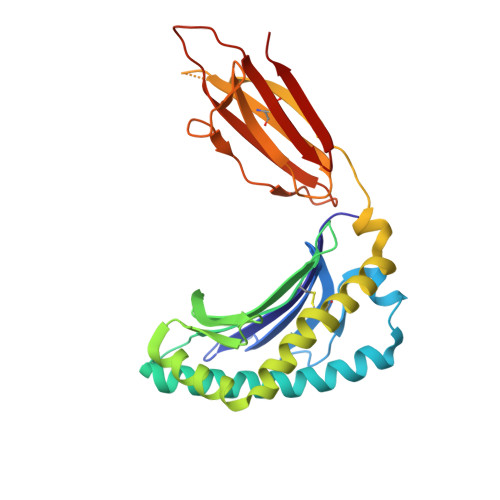
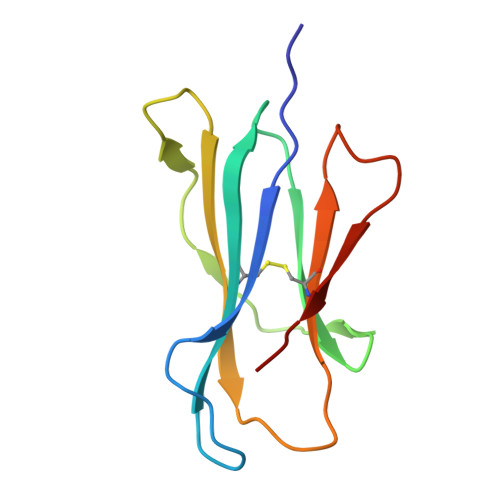
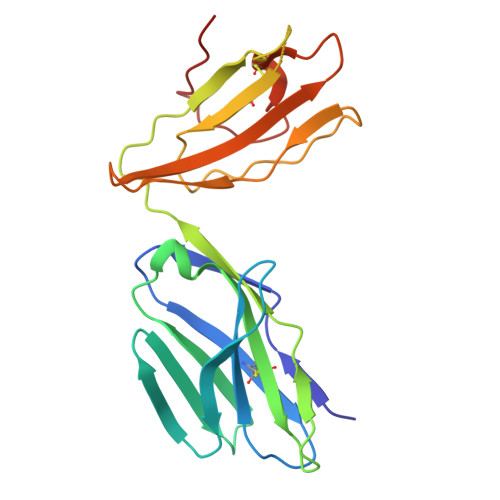
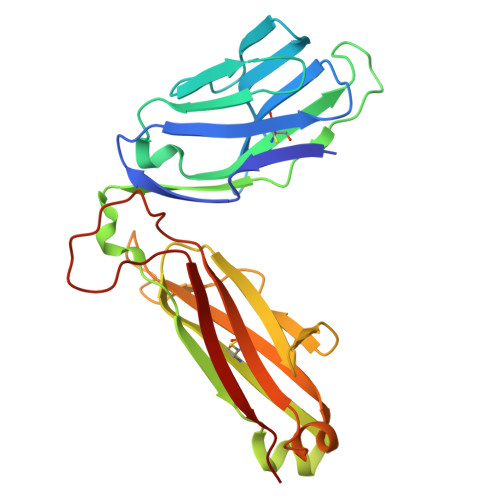
MR1 presents vitamin B6-related compounds for recognition by MR1-reactive T cells.
McInerney, M.P., Awad, W., Souter, M.N.T., Kang, Y., Wang, C.J.H., Chan Yew Poa, K., Abdelaal, M.R., Le, N.H., Shepherd, C.M., McNeice, C., Meehan, L.J., Nelson, A.G., Raynes, J.M., Mak, J.Y.W., McCluskey, J., Chen, Z., Ang, C.S., Fairlie, D.P., Le Nours, J., Illing, P.T., Rossjohn, J., Purcell, A.W.(2024) Proc Natl Acad Sci U S A 121: e2414792121-e2414792121
- PubMed: 39589872
- DOI: https://doi.org/10.1073/pnas.2414792121
- Primary Citation of Related Structures:
9CGR, 9CGS - PubMed Abstract:
The major histocompatibility complex class I related protein (MR1) presents microbially derived vitamin B2 precursors to mucosal-associated invariant T (MAIT) cells. MR1 can also present other metabolites to activate MR1-restricted T cells expressing more diverse T cell receptors (TCRs), some with anti-tumor reactivity. However, knowledge of the range of the antigen(s) that can activate diverse MR1-reactive T cells remains incomplete. Here, we identify pyridoxal (vitamin B6) as a naturally presented MR1 ligand using unbiased mass spectrometry analyses of MR1-bound metabolites. Pyridoxal, and the related compound, pyridoxal 5-phosphate bound to MR1 and enabled cell surface upregulation of wild type MR1*01 and MR1 expressing the Arg9His polymorphism associated with the MR1*04 allotype in a manner dependent on Lys43-mediated Schiff-base formation. Crystal structures of MR1*01 in complex with pyridoxal and pyridoxal 5-phosphate showed how these ligands were accommodated within the A-pocket of MR1. T cell lines transduced with the 7.G5 TCR, which has reported "pan-cancer" specificity, were specifically activated by pyridoxal presented by antigen-presenting cells expressing MR1*01 and MR1 allotypes bearing the less common Arg9His polymorphism. 7.G5 T cells also recognized, to a lesser extent, pyridoxal 5-phosphate and, importantly, recognition of both vitamers was blocked by an anti-MR1 antibody. 7.G5 TCR reactivity toward pyridoxal was enhanced when presented by the Arg9His polymorphism-bearing MR1 allotypes. Vitamin B6, and vitamers thereof, have been associated with various cancers, and here we describe a link between this ligand, MR1, and its allomorphs, and the pan-cancer 7.G5 TCR. This work identifies an MR1 ligand that can activate a diverse MR1-restricted TCR.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology and Immunity Program, Biomedicine Discovery Institute, Monash University, Clayton, VIC 3800, Australia.