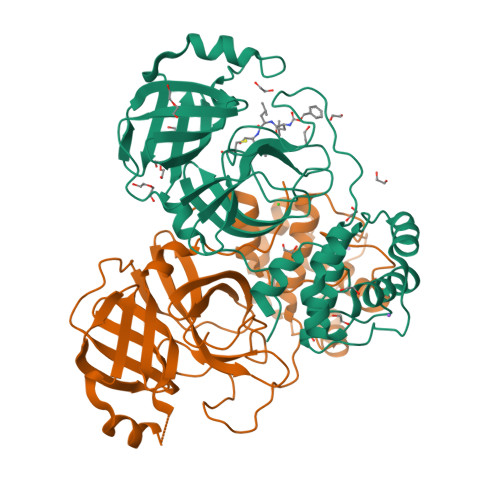
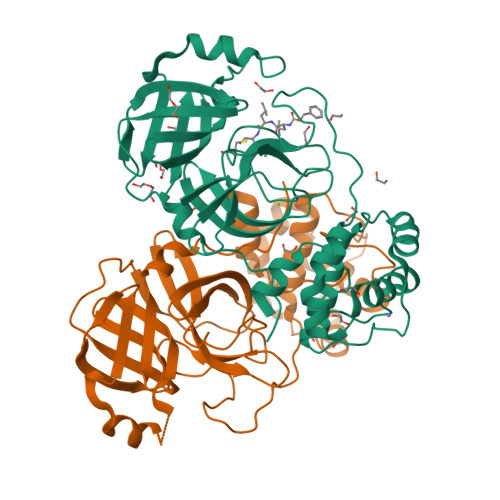
Unexpected Single-Ligand Occupancy and Negative Cooperativity in the SARS-CoV-2 Main Protease.
Albani, S., Costanzi, E., Hoang, G.L., Kuzikov, M., Frings, M., Ansari, N., Demitri, N., Nguyen, T.T., Rizzi, V., Schulz, J.B., Bolm, C., Zaliani, A., Carloni, P., Storici, P., Rossetti, G.(2024) J Chem Inf Model 64: 892-904
- PubMed: 38051605
- DOI: https://doi.org/10.1021/acs.jcim.3c01497
- Primary Citation of Related Structures:
7PHZ, 8P54, 8P55, 8P56, 8P57, 8P58, 8P5A, 8P5B, 8P5C, 8P86, 8P87 - PubMed Abstract:
Many homodimeric enzymes tune their functions by exploiting either negative or positive cooperativity between subunits. In the SARS-CoV-2 Main protease (Mpro) homodimer, the latter has been suggested by symmetry in most of the 500 reported protease/ligand complex structures solved by macromolecular crystallography (MX). Here we apply the latter to both covalent and noncovalent ligands in complex with Mpro. Strikingly, our experiments show that the occupation of both active sites of the dimer originates from an excess of ligands. Indeed, cocrystals obtained using a 1:1 ligand/protomer stoichiometry lead to single occupation only. The empty binding site exhibits a catalytically inactive geometry in solution, as suggested by molecular dynamics simulations. Thus, Mpro operates through negative cooperativity with the asymmetric activity of the catalytic sites. This allows it to function with a wide range of substrate concentrations, making it resistant to saturation and potentially difficult to shut down, all properties advantageous for the virus' adaptability and resistance.
Organizational Affiliation:
Institute for Neuroscience and Medicine (INM-9), Forschungszentrum Jülich, Jülich 52425, Germany.