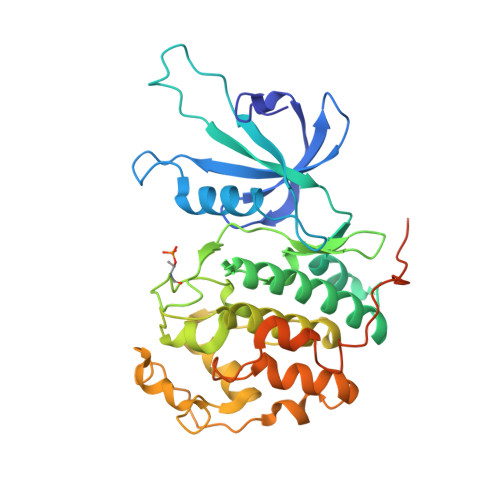
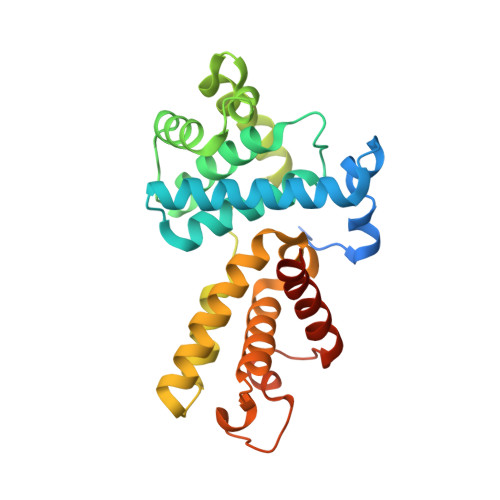
The reversible inhibitor SR-4835 binds Cdk12/cyclin K in a noncanonical G-loop conformation.
Schmitz, M., Kaltheuner, I.H., Anand, K., Duster, R., Moecking, J., Monastyrskyi, A., Duckett, D.R., Roush, W.R., Geyer, M.(2023) J Biol Chem 300: 105501-105501
- PubMed: 38016516
- DOI: https://doi.org/10.1016/j.jbc.2023.105501
- Primary Citation of Related Structures:
8P81 - PubMed Abstract:
Inhibition of cyclin-dependent kinases (CDKs) has evolved as an emerging anticancer strategy. In addition to the cell cycle-regulating CDKs, the transcriptional kinases Cdk12 and Cdk13 have become the focus of interest as they mediate a variety of functions, including the transition from transcription initiation to elongation and termination, precursor mRNA splicing, and intronic polyadenylation. Here, we determine the crystal structure of the small molecular inhibitor SR-4835 bound to the Cdk12/cyclin K complex at 2.68 Å resolution. The compound's benzimidazole moiety is embedded in a unique hydrogen bond network mediated by the kinase hinge region with flanking hydroxy groups of the Y815 and D819 side chains. Whereas the SR-4835 head group targets the adenine-binding pocket, the kinase's glycine-rich loop is shifted down toward the activation loop. Additionally, the αC-helix adopts an inward conformation, and the phosphorylated T-loop threonine interacts with all three canonical arginines, a hallmark of CDK activation that is altered in Cdk12 and Cdk13. Dose-response inhibition measurements with recombinant CMGC kinases show that SR-4835 is highly specific for Cdk12 and Cdk13 following a 10-fold lower potency for Cdk10. Whereas other CDK-targeting compounds exhibit tighter binding affinities and higher potencies for kinase inhibition, SR-4835 can be considered a selective transcription elongation antagonist. Our results provide the basis for a rational improvement of SR-4835 toward Cdk12 inhibition and a gain in selectivity over other transcription regulating CDKs.
Organizational Affiliation:
Institute of Structural Biology, University of Bonn, Bonn, Germany.