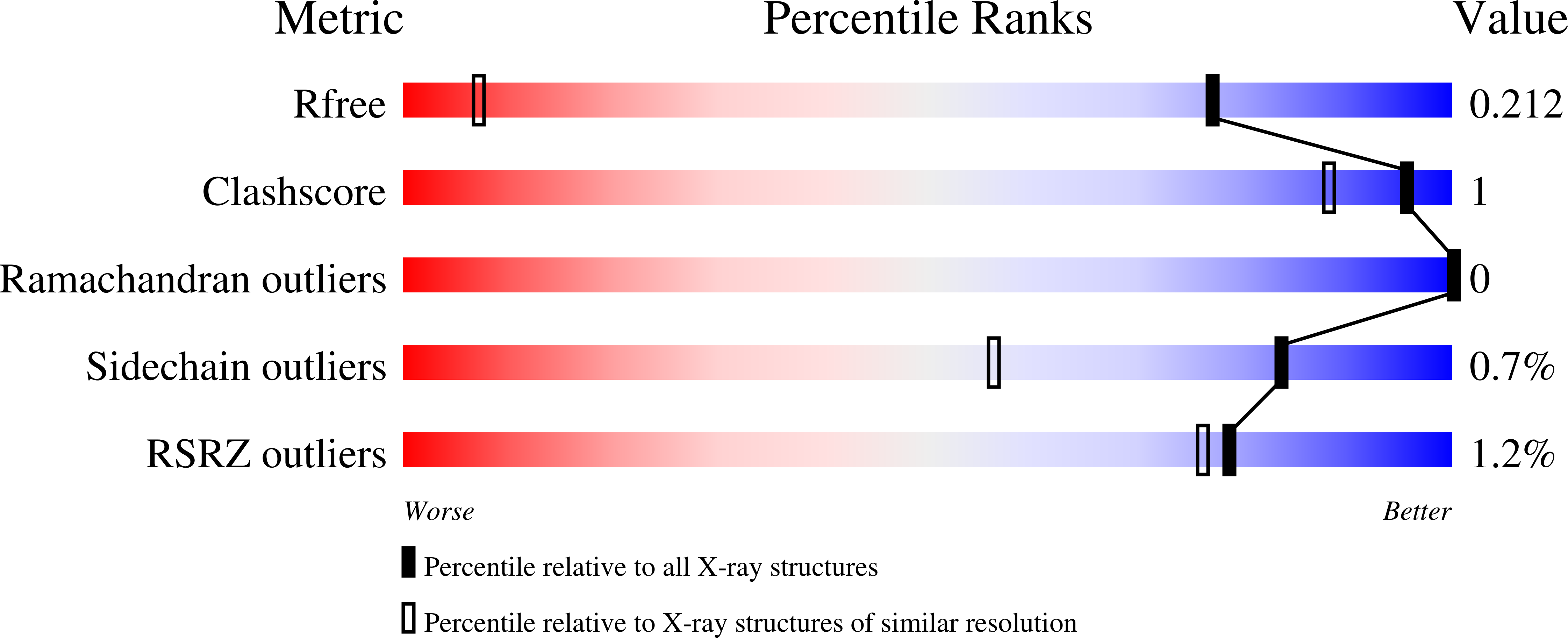
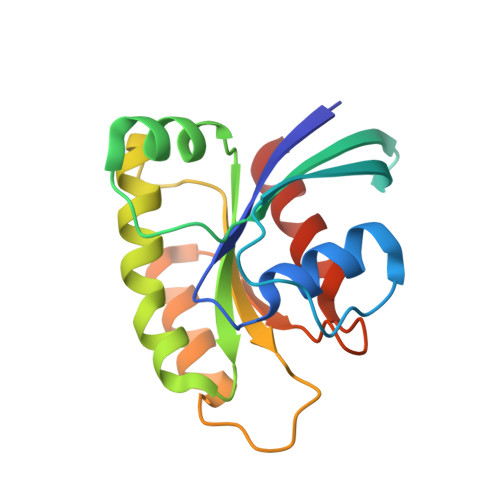
Fragment Optimization of Reversible Binding to the Switch II Pocket on KRAS Leads to a Potent, In Vivo Active KRAS G12C Inhibitor.
Broker, J., Waterson, A.G., Smethurst, C., Kessler, D., Bottcher, J., Mayer, M., Gmaschitz, G., Phan, J., Little, A., Abbott, J.R., Sun, Q., Gmachl, M., Rudolph, D., Arnhof, H., Rumpel, K., Savarese, F., Gerstberger, T., Mischerikow, N., Treu, M., Herdeis, L., Wunberg, T., Gollner, A., Weinstabl, H., Mantoulidis, A., Kramer, O., McConnell, D.B., W Fesik, S.(2022) J Med Chem 65: 14614-14629
- PubMed: 36300829
- DOI: https://doi.org/10.1021/acs.jmedchem.2c01120
- Primary Citation of Related Structures:
7U8H, 8AFB, 8AFC, 8AFD - PubMed Abstract:
Activating mutations in KRAS are the most frequent oncogenic alterations in cancer. The oncogenic hotspot position 12, located at the lip of the switch II pocket, offers a covalent attachment point for KRAS G12C inhibitors. To date, KRAS G12C inhibitors have been discovered by first covalently binding to the cysteine at position 12 and then optimizing pocket binding. We report on the discovery of the in vivo active KRAS G12C inhibitor BI-0474 using a different approach, in which small molecules that bind reversibly to the switch II pocket were identified and then optimized for non-covalent binding using structure-based design. Finally, the Michael acceptor containing warhead was attached. Our approach offers not only an alternative approach to discovering KRAS G12C inhibitors but also provides a starting point for the discovery of inhibitors against other oncogenic KRAS mutants.
Organizational Affiliation:
Boehringer Ingelheim RCV GmbH & Co. KG, Dr. Boehringer Gasse 5-11, A-1121 Vienna, Austria.