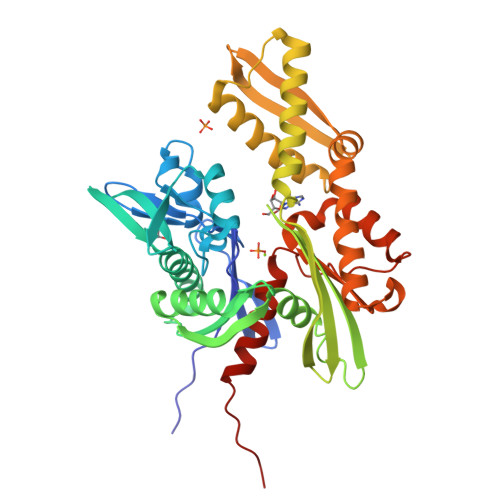
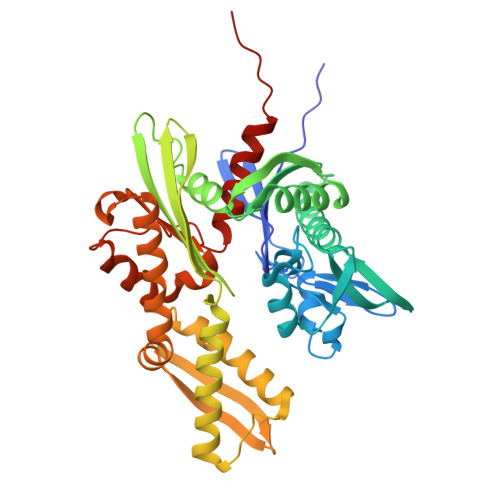
Specific radiation damage to halogenated inhibitors and ligands in protein-ligand crystal structures.
Rodrigues, M.J., Cabry, M., Collie, G., Carter, M., McAndrew, C., Owen, R.L., Bellenie, B.R., Le Bihan, Y.V., van Montfort, R.L.M.(2024) J Appl Crystallogr 57: 1951-1965
- PubMed: 39628887
- DOI: https://doi.org/10.1107/S1600576724010549
- Primary Citation of Related Structures:
7GUD, 7GUE, 7GUF, 7GUG, 7GUH, 7GUI, 7GUJ, 7GUK, 7GUL, 7GUM, 7GUN, 7GUO, 7GUP, 7GUQ, 7GUR, 7GUS, 7GUT, 7GUU, 7GUV, 7GUW, 7GUX, 7GUY, 7GUZ, 7GV0, 7GV1, 7GV2, 7GV3, 7GV4, 7GV5, 7GV6, 7GV7, 7GV8, 7GV9, 7GVA, 7GVB, 7GVC, 7GVD, 7GVE, 7GVF, 7GVG, 7GVH, 7GVI, 7GVJ, 7GVK, 7GVL, 7GVM, 7GVN, 7GVO, 7GVP, 7GVQ - PubMed Abstract:
Protein-inhibitor crystal structures aid medicinal chemists in efficiently improving the potency and selectivity of small-molecule inhibitors. It is estimated that a quarter of lead molecules in drug discovery projects are halogenated. Protein-inhibitor crystal structures have shed light on the role of halogen atoms in ligand binding. They form halogen bonds with protein atoms and improve shape complementarity of inhibitors with protein binding sites. However, specific radiation damage (SRD) can cause cleavage of carbon-halogen (C- X ) bonds during X-ray diffraction data collection. This study shows significant C- X bond cleavage in protein-ligand structures of the therapeutic cancer targets B-cell lymphoma 6 (BCL6) and heat shock protein 72 (HSP72) complexed with halogenated ligands, which is dependent on the type of halogen and chemical structure of the ligand. The study found that metrics used to evaluate the fit of the ligand to the electron density deteriorated with increasing X-ray dose, and that SRD eliminated the anomalous signal from brominated ligands. A point of diminishing returns is identified, where collecting highly redundant data reduces the anomalous signal that may be used to identify binding sites of low-affinity ligands or for experimental phasing. Straightforward steps are proposed to mitigate the effects of C- X bond cleavage on structures of proteins bound to halogenated ligands and to improve the success of anomalous scattering experiments.
Organizational Affiliation:
Centre for Cancer Drug Discovery The Institute of Cancer Research 15 Cotswold Road Sutton LondonSM2 5NG United Kingdom.