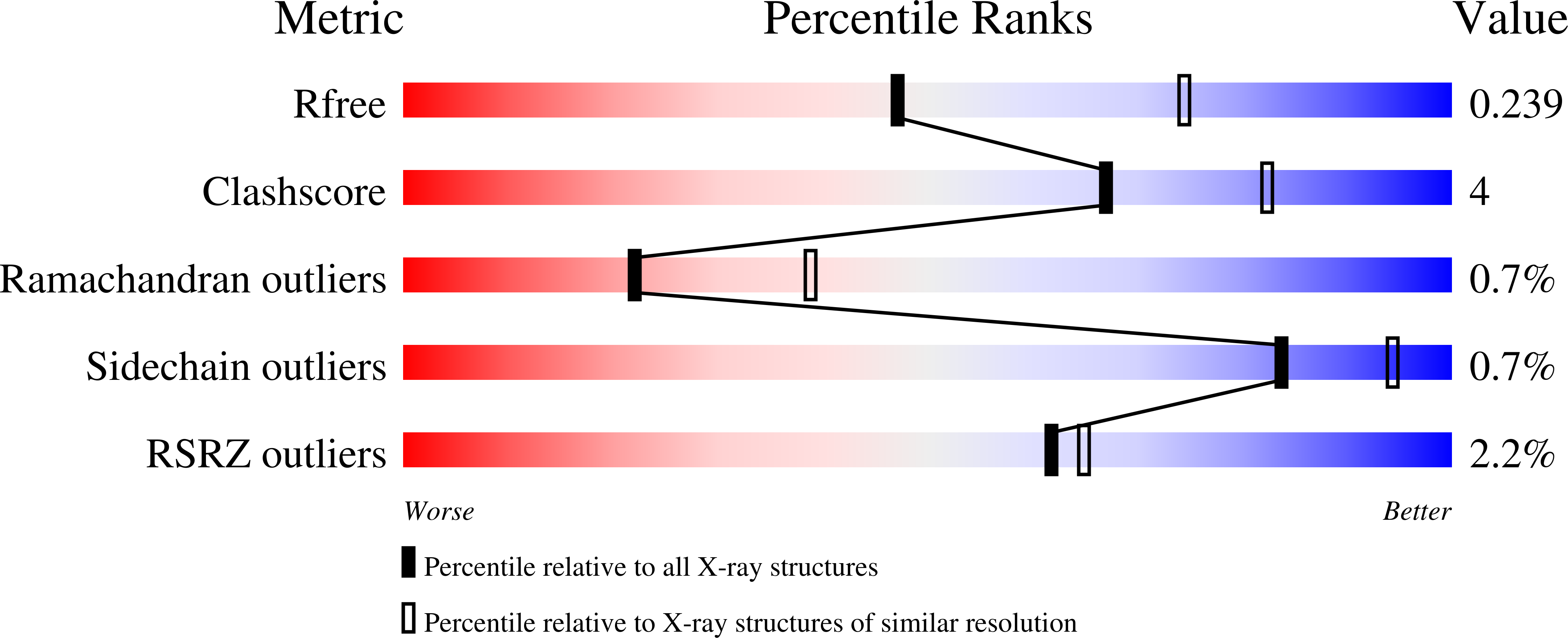
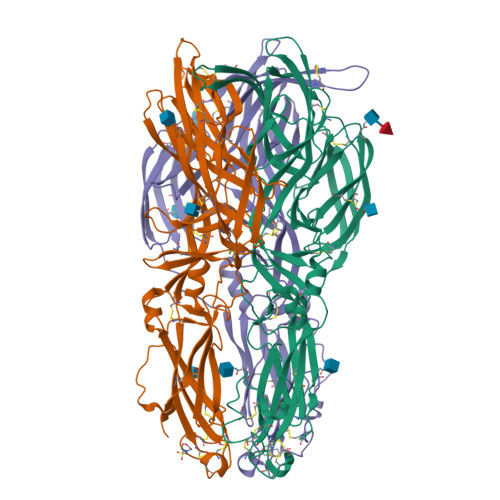
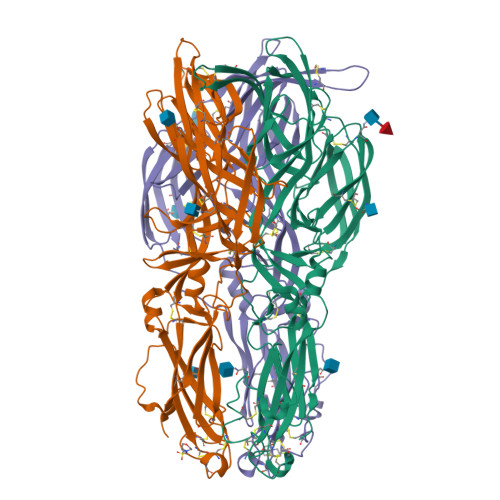
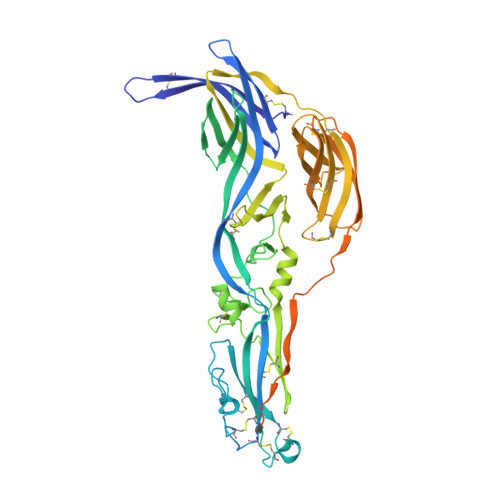
A glycerophospholipid-specific pocket in the RVFV class II fusion protein drives target membrane insertion.
Guardado-Calvo, P., Atkovska, K., Jeffers, S.A., Grau, N., Backovic, M., Perez-Vargas, J., de Boer, S.M., Tortorici, M.A., Pehau-Arnaudet, G., Lepault, J., England, P., Rottier, P.J., Bosch, B.J., Hub, J.S., Rey, F.A.(2017) Science 358: 663-667
- PubMed: 29097548
- DOI: https://doi.org/10.1126/science.aal2712
- Primary Citation of Related Structures:
6EGT, 6EGU - PubMed Abstract:
The Rift Valley fever virus (RVFV) is transmitted by infected mosquitoes, causing severe disease in humans and livestock across Africa. We determined the x-ray structure of the RVFV class II fusion protein Gc in its postfusion form and in complex with a glycerophospholipid (GPL) bound in a conserved cavity next to the fusion loop. Site-directed mutagenesis and molecular dynamics simulations further revealed a built-in motif allowing en bloc insertion of the fusion loop into membranes, making few nonpolar side-chain interactions with the aliphatic moiety and multiple polar interactions with lipid head groups upon membrane restructuring. The GPL head-group recognition pocket is conserved in the fusion proteins of other arthropod-borne viruses, such as Zika and chikungunya viruses, which have recently caused major epidemics worldwide.
Organizational Affiliation:
Institut Pasteur, Département de Virologie, Unité de Virologie Structurale, 75724 Paris Cedex 15, France. pablo.guardado-calvo@pasteur.fr jhub@gwdg.de felix.rey@pasteur.fr.