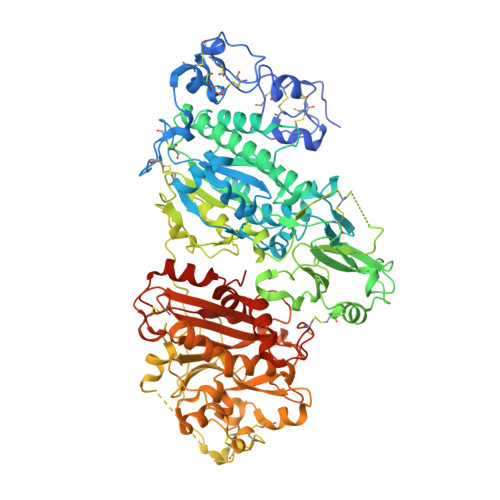
Discovery of BI-2545: A Novel Autotaxin Inhibitor That Significantly Reduces LPA Levels in Vivo.
Kuttruff, C.A., Ferrara, M., Bretschneider, T., Hoerer, S., Handschuh, S., Nosse, B., Romig, H., Nicklin, P., Roth, G.J.(2017) ACS Med Chem Lett 8: 1252-1257
- PubMed: 29259743
- DOI: https://doi.org/10.1021/acsmedchemlett.7b00312
- Primary Citation of Related Structures:
5OHI, 5OLB - PubMed Abstract:
In an effort to find new therapeutic interventions addressing the unmet medical need of patients with idiopathic pulmonary fibrosis, we initiated a program to identify new autotaxin (ATX) inhibitors. Starting from a recently published compound (PF-8380), we identified several highly potent ATX inhibitors with improved pharmacokinetic and safety profiles. Further optimization efforts resulted in the identification of a single-digit nanomolar lead compound (BI-2545) that shows substantial lowering of LPA in vivo and is therefore considered a valuable tool for further studies.
Organizational Affiliation:
Medicinal Chemistry, Drug Discovery Sciences, and Immunology & Respiratory, Boehringer Ingelheim Pharma GmbH & Co. KG, 88397 Biberach an der Riss, Germany.