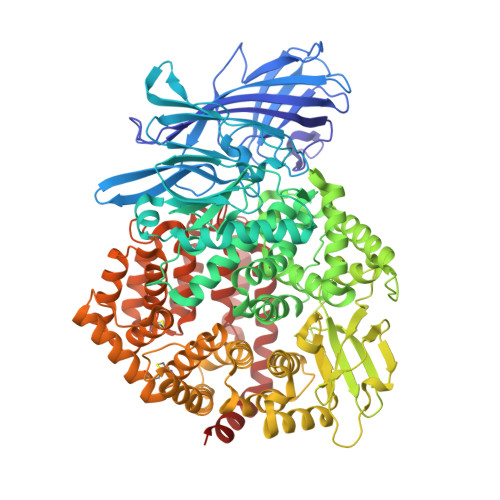
The X-ray Crystal Structure of Human Aminopeptidase N Reveals a Novel Dimer and the Basis for Peptide Processing.
Wong, A.H., Zhou, D., Rini, J.M.(2012) J Biol Chem 287: 36804-36813
- PubMed: 22932899
- DOI: https://doi.org/10.1074/jbc.M112.398842
- Primary Citation of Related Structures:
4FYQ, 4FYR, 4FYS, 4FYT - PubMed Abstract:
Human aminopeptidase N (hAPN/hCD13) is a dimeric membrane protein and a member of the M1 family of zinc metallopeptidases. Within the rennin-angiotensin system, its enzymatic activity is responsible for processing peptide hormones angiotensin III and IV. In addition, hAPN is also involved in cell adhesion, endocytosis, and signal transduction and it is an important target for cancer therapy. Reported here are the high resolution x-ray crystal structures of the dimeric ectodomain of hAPN and its complexes with angiotensin IV and the peptidomimetic inhibitors, amastatin and bestatin. Each monomer of the dimer is found in what has been termed the closed form in other M1 enzymes and each monomer is characterized by an internal cavity surrounding the catalytic site as well as a unique substrate/inhibitor-dependent loop ordering, which in the case of the bestatin complex suggests a new route to inhibitor design. The hAPN structure provides the first example of a dimeric M1 family member and the observed structural features, in conjunction with a model for the open form, provide novel insights into the mechanism of peptide processing and signal transduction.
Organizational Affiliation:
Department of Biochemistry, University of Toronto, Toronto, Ontario M5S 1A8, Canada.