Pharmacological chaperones for human alpha-N-acetylgalactosaminidase
Clark, N.E., Metcalf, M.C., Best, D., Fleet, G.W., Garman, S.C.(2012) Proc Natl Acad Sci U S A 109: 17400-17405
- PubMed: 23045655
- DOI: https://doi.org/10.1073/pnas.1203924109
- Primary Citation of Related Structures:
4DO4, 4DO5, 4DO6 - PubMed Abstract:
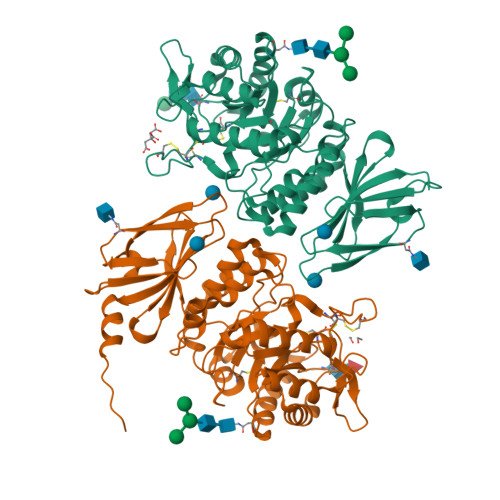
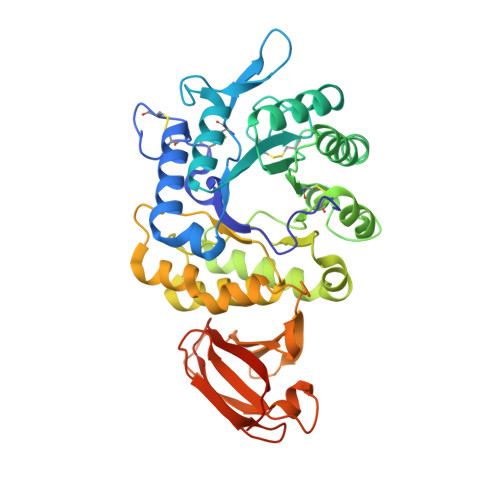
Schindler/Kanzaki disease is an inherited metabolic disease with no current treatment options. This neurologic disease results from a defect in the lysosomal α-N-acetylgalactosaminidase (α-NAGAL) enzyme. In this report, we show evidence that the iminosugar DGJNAc can inhibit, stabilize, and chaperone human α-NAGAL both in vitro and in vivo. We demonstrate that a related iminosugar DGJ (currently in phase III clinical trials for another metabolic disorder, Fabry disease) can also chaperone human α-NAGAL in Schindler/Kanzaki disease. The 1.4- and 1.5-Å crystal structures of human α-NAGAL complexes reveal the different binding modes of iminosugars compared with glycosides. We show how differences in two functional groups result in >9 kcal/mol of additional binding energy and explain the molecular interactions responsible for the unexpectedly high affinity of the pharmacological chaperones. These results open two avenues for treatment of Schindler/Kanzaki disease and elucidate the atomic basis for pharmacological chaperoning in the entire family of lysosomal storage diseases.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of Massachusetts, Amherst, MA 01003, USA.