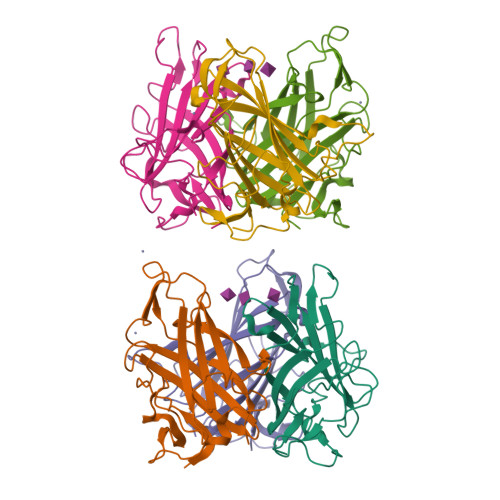
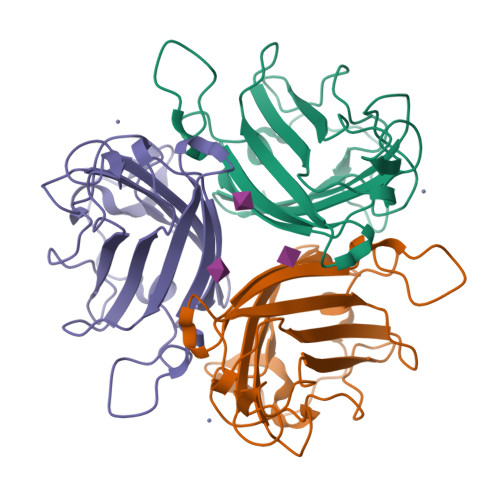
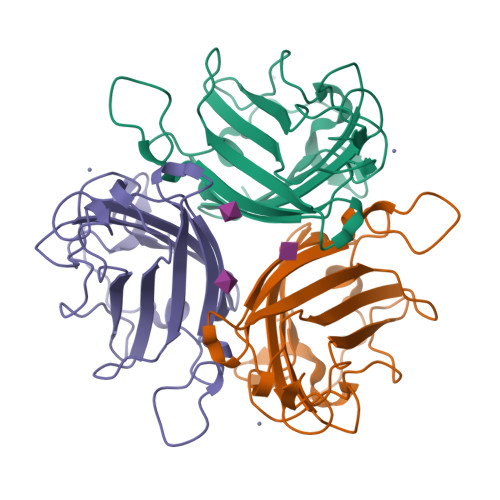
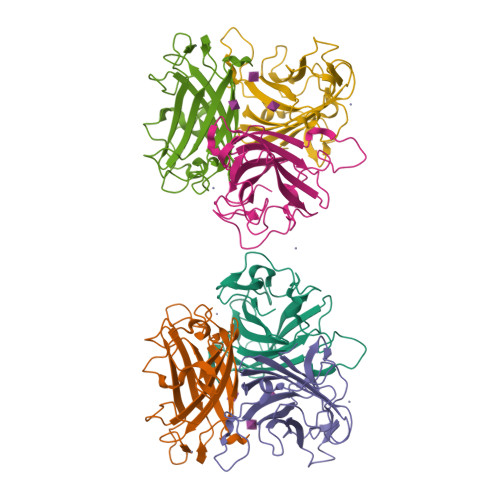
A potent trivalent sialic Acid inhibitor of adenovirus type 37 infection of human corneal cells.
Spjut, S., Qian, W., Bauer, J., Storm, R., Frangsmyr, L., Stehle, T., Arnberg, N., Elofsson, M.(2011) Angew Chem Int Ed Engl 50: 6519-6521
- PubMed: 21648036
- DOI: https://doi.org/10.1002/anie.201101559
- Primary Citation of Related Structures:
3QND
Organizational Affiliation:
Department of Chemistry, Umeå Centre for Microbial Research (UCMR) and Laboratory for Molecular Infection Medicine Sweden (MIMS), Umeå University, 90187 Umeå, Sweden.