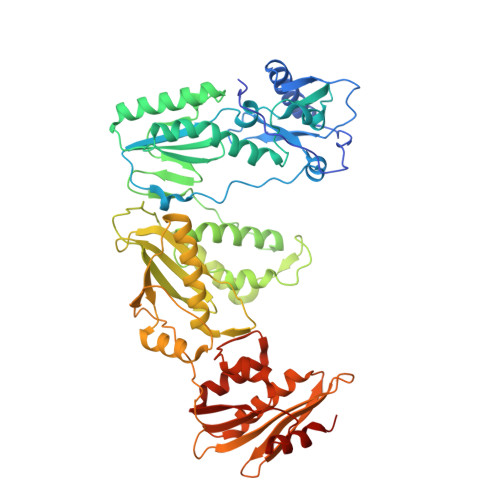
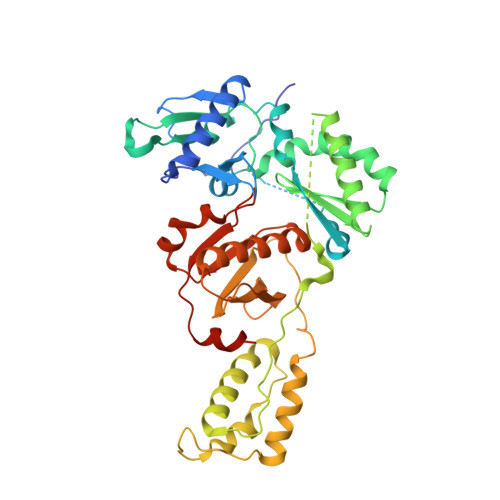
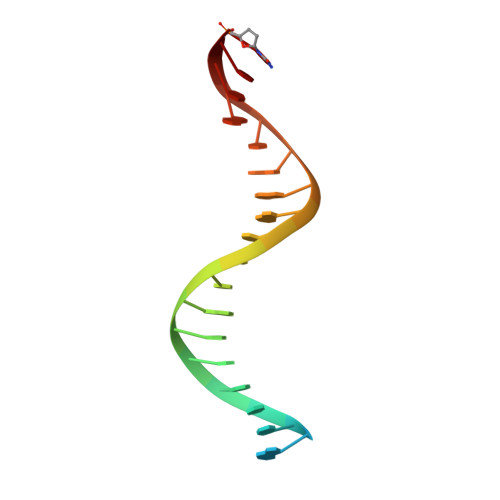
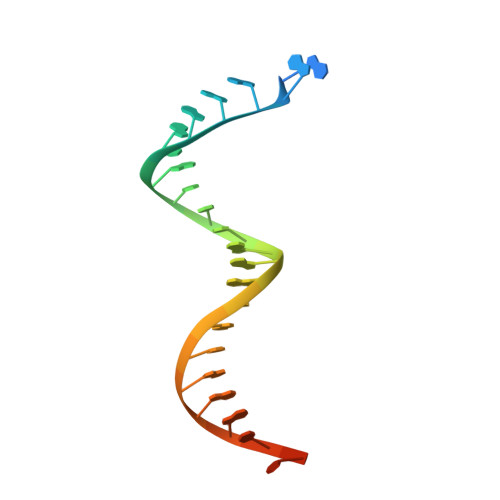
Elucidating molecular interactions ofL-nucleotides with HIV-1 reverse transcriptase and mechanism of M184V-caused drug resistance.
Hung, M., Tokarsky, E.J., Lagpacan, L., Zhang, L., Suo, Z., Lansdon, E.B.(2019) Commun Biol 2: 469-469
- PubMed: 31872074
- DOI: https://doi.org/10.1038/s42003-019-0706-x
- Primary Citation of Related Structures:
6UIR, 6UIS, 6UIT, 6UJX, 6UJY, 6UJZ, 6UK0 - PubMed Abstract:
Emtricitabine (FTC) and lamivudine (3TC), containing an oxathiolane ring with unnatural (-)-stereochemistry, are widely used nucleoside reverse transcriptase inhibitors (NRTIs) in anti-HIV therapy. Treatment with FTC or 3TC primarily selects for the HIV-1 RT M184V/I resistance mutations. Here we provide a comprehensive kinetic and structural basis for inhibiting HIV-1 RT by (-)-FTC-TP and (-)-3TC-TP and drug resistance by M184V. (-)-FTC-TP and (-)-3TC-TP have higher binding affinities (1/ K d ) for wild-type RT but slower incorporation rates than dCTP. HIV-1 RT ternary crystal structures with (-)-FTC-TP and (-)-3TC-TP corroborate kinetic results demonstrating that their oxathiolane sulfur orients toward the DNA primer 3'-terminus and their triphosphate exists in two different binding conformations. M184V RT displays greater (>200-fold) K d for the L -nucleotides and moderately higher (>9-fold) K d for the D -isomers compared to dCTP. The M184V RT structure illustrates how the mutation repositions the oxathiolane of (-)-FTC-TP and shifts its triphosphate into a non-productive conformation.
Organizational Affiliation:
1Gilead Sciences, Inc., 333 Lakeside Dr., Foster City, CA 94404 USA.