Advances in long-wavelength native phasing at X-ray free-electron lasers.
Nass, K., Cheng, R., Vera, L., Mozzanica, A., Redford, S., Ozerov, D., Basu, S., James, D., Knopp, G., Cirelli, C., Martiel, I., Casadei, C., Weinert, T., Nogly, P., Skopintsev, P., Usov, I., Leonarski, F., Geng, T., Rappas, M., Dore, A.S., Cooke, R., Nasrollahi Shirazi, S., Dworkowski, F., Sharpe, M., Olieric, N., Bacellar, C., Bohinc, R., Steinmetz, M.O., Schertler, G., Abela, R., Patthey, L., Schmitt, B., Hennig, M., Standfuss, J., Wang, M., Milne, C.J.(2020) IUCrJ 7: 965-975
- PubMed: 33209311
- DOI: https://doi.org/10.1107/S2052252520011379
- Primary Citation of Related Structures:
6S0L, 6S0Q, 6S19, 6S1D, 6S1E, 6S1G - PubMed Abstract:
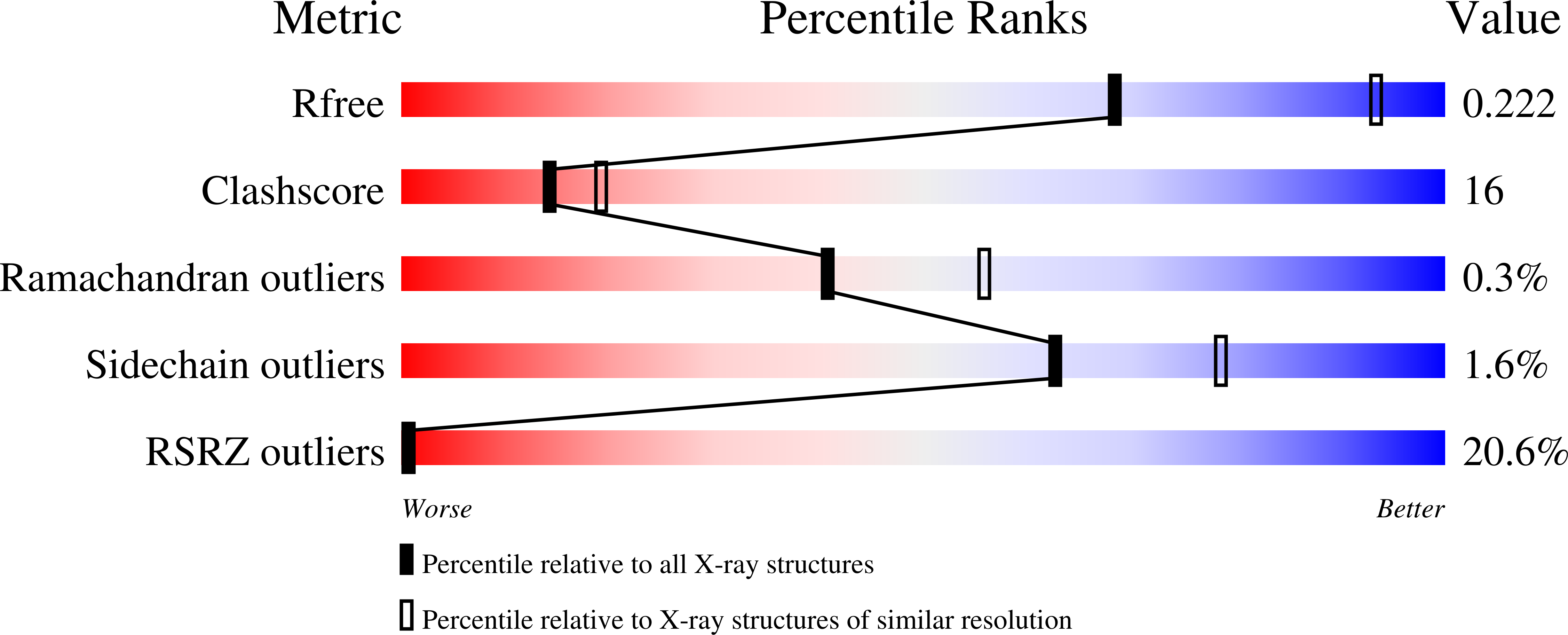
Long-wavelength pulses from the Swiss X-ray free-electron laser (XFEL) have been used for de novo protein structure determination by native single-wavelength anomalous diffraction (native-SAD) phasing of serial femtosecond crystallography (SFX) data. In this work, sensitive anomalous data-quality indicators and model proteins were used to quantify improvements in native-SAD at XFELs such as utilization of longer wavelengths, careful experimental geometry optimization, and better post-refinement and partiality correction. Compared with studies using shorter wavelengths at other XFELs and older software versions, up to one order of magnitude reduction in the required number of indexed images for native-SAD was achieved, hence lowering sample consumption and beam-time requirements significantly. Improved data quality and higher anomalous signal facilitate so-far underutilized de novo structure determination of challenging proteins at XFELs. Improvements presented in this work can be used in other types of SFX experiments that require accurate measurements of weak signals, for example time-resolved studies.
Organizational Affiliation:
Photon Science Division, Paul Scherrer Institut, Forschungsstrasse 111, Villigen PSI, 5232, Switzerland.