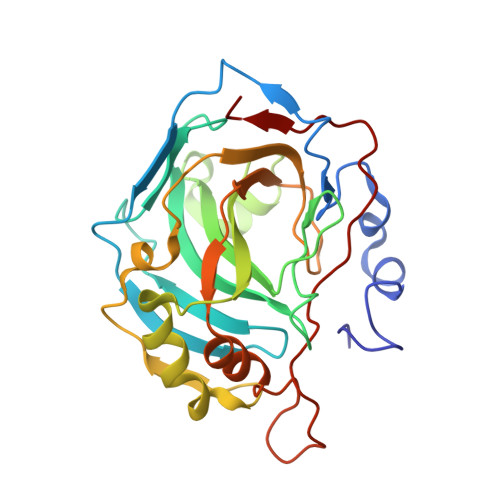
Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Bhatt, A., Mahon, B.P., Cruzeiro, V.W., Cornelio, B., Laronze-Cochard, M., Ceruso, M., Sapi, J., Rance, G.A., Khlobystov, A.N., Fontana, A., Roitberg, A., Supuran, C.T., McKenna, R.(2017) Chembiochem 18: 213-222
- PubMed: 27860128
- DOI: https://doi.org/10.1002/cbic.201600513
- Primary Citation of Related Structures:
5SZ0, 5SZ1, 5SZ2, 5SZ3, 5SZ4, 5SZ5, 5SZ6, 5SZ7 - PubMed Abstract:
Carbonic anhydrases (CAs) are implicated in a wide range of diseases, including the upregulation of isoforms CA IX and XII in many aggressive cancers. However, effective inhibition of disease-implicated CAs should minimally affect the ubiquitously expressed isoforms, including CA I and II, to improve directed distribution of the inhibitors to the cancer-associated isoforms and reduce side effects. Four benzenesulfonamide-based inhibitors were synthesized by using the tail approach and displayed nanomolar affinities for several CA isoforms. The crystal structures of the inhibitors bound to a CA IX mimic and CA II are presented. Further in silico modeling was performed with the inhibitors docked into CA I and XII to identify residues that contributed to or hindered their binding interactions. These structural studies demonstrated that active-site residues lining the hydrophobic pocket, especially positions 92 and 131, dictate the positional binding and affinity of inhibitors, whereas the tail groups modulate CA isoform specificity. Geometry optimizations were performed on each ligand in the crystal structures and showed that the energetic penalties of the inhibitor conformations were negligible compared to the gains from active-site interactions. These studies further our understanding of obtaining isoform specificity when designing small molecule CA inhibitors.
Organizational Affiliation:
Department of, Biochemistry and Molecular Biology, College of Medicine, University of Florida, P. O. Box 100245, Gainesville, FL, 32610, USA.