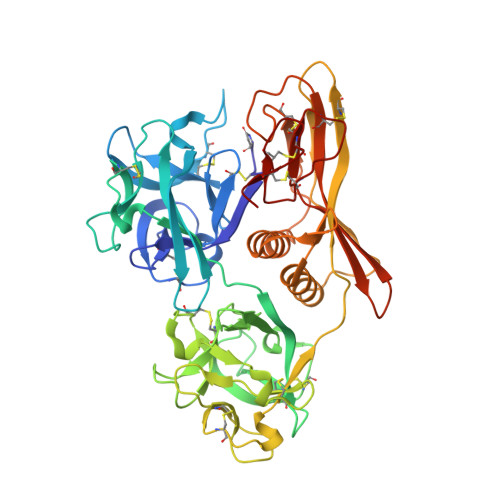
Crystal Structure of the Hemolytic Lectin CEL-III Isolated from the Marine Invertebrate Cucumaria echinata: IMPLICATIONS OF DOMAIN STRUCTURE FOR ITS MEMBRANE PORE-FORMATION MECHANISM
Uchida, T., Yamasaki, T., Eto, S., Sugawara, H., Kurisu, G., Nakagawa, A., Kusunoki, M., Hatakeyama, T.(2004) J Biol Chem 279: 37133-37141
- PubMed: 15194688
- DOI: https://doi.org/10.1074/jbc.M404065200
- Primary Citation of Related Structures:
1VCL - PubMed Abstract:
CEL-III is a Ca(2+)-dependent and galactose-specific lectin purified from the sea cucumber, Cucumaria echinata, which exhibits hemolytic and hemagglutinating activities. Six molecules of CEL-III are assumed to oligomerize to form an ion-permeable pore in the cell membrane. We have determined the crystal structure of CELIII by using single isomorphous replacement aided by anomalous scattering in lead at 1.7 A resolution. CEL-III consists of three distinct domains as follows: the N-terminal two carbohydrate-binding domains (1 and 2), which adopt beta-trefoil folds such as the B-chain of ricin and are members of the (QXW)(3) motif family; and domain 3, which is a novel fold composed of two alpha-helices and one beta-sandwich. CEL-III is the first Ca(2+)-dependent lectin structure with two beta-trefoil folds. Despite sharing the structure of the B-chain of ricin, CEL-III binds five Ca(2+) ions at five of the six subdomains in both domains 1 and 2. Considering the relatively high similarity among the five subdomains, they are putative binding sites for galactose-related carbohydrates, although it remains to be elucidated whether bound Ca(2+) is directly involved in interaction with carbohydrates. The paucity of hydrophobic interactions in the interfaces between the domains and biochemical data suggest that these domains rearrange upon carbohydrate binding in the erythrocyte membrane. This conformational change may be responsible for oligomerization of CEL-III molecules and hemolysis in the erythrocyte membranes.
Organizational Affiliation:
Research Center for Structural and Functional Proteomics, Institute for Protein Research, Osaka University, 3-2 Yamadaoka, Suita, Osaka 565-0871.