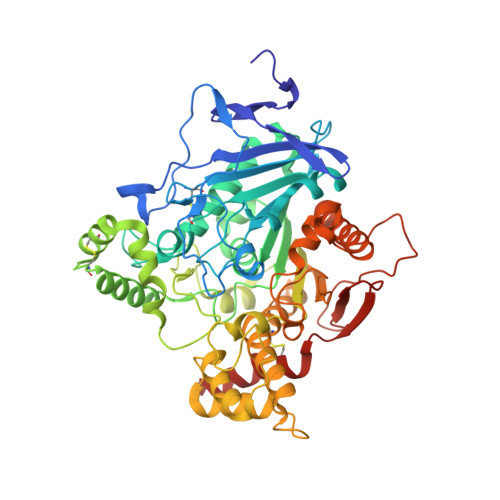
Structural insights into ligand interactions at the acetylcholinesterase peripheral anionic site
Bourne, Y., Taylor, P., Radic, Z., Marchot, P.(2003) EMBO J 22: 1-12
- PubMed: 12505979
- DOI: https://doi.org/10.1093/emboj/cdg005
- Primary Citation of Related Structures:
1J06, 1J07, 1KU6, 1N5M, 1N5R - PubMed Abstract:
The peripheral anionic site on acetylcholinesterase (AChE), located at the active center gorge entry, encompasses overlapping binding sites for allosteric activators and inhibitors; yet, the molecular mechanisms coupling this site to the active center at the gorge base to modulate catalysis remain unclear. The peripheral site has also been proposed to be involved in heterologous protein associations occurring during synaptogenesis or upon neurodegeneration. A novel crystal form of mouse AChE, combined with spectrophotometric analyses of the crystals, enabled us to solve unique structures of AChE with a free peripheral site, and as three complexes with peripheral site inhibitors: the phenylphenanthridinium ligands, decidium and propidium, and the pyrogallol ligand, gallamine, at 2.20-2.35 A resolution. Comparison with structures of AChE complexes with the peptide fasciculin or with organic bifunctional inhibitors unveils new structural determinants contributing to ligand interactions at the peripheral site, and permits a detailed topographic delineation of this site. Hence, these structures provide templates for designing compounds directed to the enzyme surface that modulate specific surface interactions controlling catalytic activity and non-catalytic heterologous protein associations.
Organizational Affiliation:
Architecture et Fonction des Macromolécules Biologiques, CNRS UMR 6098, Marseille, France. [email protected]